Gluten Be Gone: Synthetic Biology Solution for Celiac Disease
What is Celiac Disease? Celiac disease or gluten allergy comes from eating wheat, rye or barley. Most common in people of N. European descent, the symptoms include diarrhea, weight loss and an increased risk of cancer.
Why is gluten allergenic? Gluten contains an unusual protein called alpha gliadin, which has many repeats of the amino acids Proline and Glutamine (PQ motifs) that are resistant to the digestive enzymes in our stomach. In some people, these PQ-rich fragments cause severe allergy and inflammation.
Clinical trials: A natural bacterial enzyme from Sphingomonas capsulata that can break down PQ motifs is in clinical trials as an Oral Enzyme Therapeutic. But it works poorly in the acidic compartment of our stomach, and attempts to engineer it to become acid tolerant have not worked.
Trial by Acid: Univ. Washington undergraduates tackled the problem from the opposite direction. They found an enzyme called Kumamolysin-AS in a heat and acid loving bacterium Alicyclobacillus sendaiensis that was already acid tolerant. They tinkered with it, using the Fold-It protein folding game, until they found variants predicted to change the enzyme’s preference from Proline Arginine (PR) to Proline Glutamine (PQ). When they made and tested ~260 engineered enzymes, they found one that had a 116-fold increase in ability to digest the gluten peptide in acidic conditions, with a switch in preference of 800-fold! The new enzyme, KumaMAX, could be used in oral therapy or engineered into common bacteria found in yogurt to make probiotics.
So Much Win!: This work (1) could help millions of gluten allergy sufferers world wide, (2) was done by undergraduates competing in iGEM, an annual synthetic biology competition originally founded at MIT, (3) using gaming software, (4) built on basic research done on an obscure bacterial enzyme, and (5) published with student authors in a peer-reviewed journal.
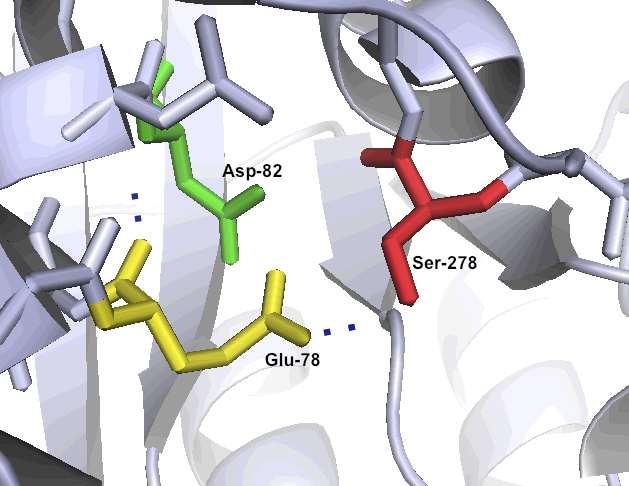
Images: Normal catalytic triad of protease enyzmes (left) and acid tolerant substitution (right) found in bacteria growing in acid, hot springs (middle).
Paper: Computational Design of an α‑Gliadin Peptidase; Gordon et al., (2012) JACS 134, 20513−20520
Team UW iGEM: http://goo.gl/vgvTX
#ScienceSunday #syntheticbiology


I think my mum’s best friend has problems with gluten. I’ll pass these details on. A ray of hope.
Funny that you post this I had them draw my blood yesterday to test for Celiac Disease. I had a food allergy test done a couple of years ago and yes gluten was something I have an allergy to. Will know here fairly soon.
In a previous thread a few weeks ago we got into considerations of changes that my have occurred in wheat quite recently that are creating celiac-like problems for a growing number of people
Thanks, Fred Gandt . I forgot to mention that the only treatment right now is a gluten-free diet, which is difficult to manage and can get expensive.
joe breskin I strongly suspect Monsanto…..
Good luck, Mark Herndon ! joe breskin is there a change in the genetic varieties of wheat grown or possibly in processing of cereals?
Interesting comparison, Dennis Bennett ! Fortunately, lactose can be digested with oral pills or by pretreating products with the enzyme lactase.
/cc: Scott Blum
http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0001280/figure/A000233.B19292/?report=objectonly One thing I have noticed is that as I have reduced out gluten whenever I can I tend not to have headaches anymore. The great thing is I see a lot of food producers making their foods gluten free.
Yes Rajini Rao. As I understand it, she has a tough time finding replacements for things or simply has to go without.
I’m just going to phone my mum and ask her to pass on this post’s address.
I’ll leave this link here, could be helpful to people with gluten sensitivity. Experts say that as much 10% of the population may show symtoms from gluten but not full blown celiac and that gluten is pretty indigestible for all of us:
http://www.cnn.com/2011/HEALTH/04/12/gluten.free.diet.improve/index.html
Mark Herndon , take a look at that CNN article I just linked to..because you may feel better with a gluten free diet but not test positive for celiac disease.
This is great news on advances to balance out the body’s inbalances to gluten.
iGEM is wonderful, I’m sure you will enjoy the experience Lucas Frib . Good luck to your team!
Well done and very informative Rajini Rao 🙂
Shaker Cherukuri , your comment prompted me to look this up. I had not realized it was this common! Apparently, “food manufacturers like using gluten as an additive in prepared foods. Gluten is used as a stablizer, an emulsifier, a thickener and flow agent in literally hundreds of processed foods, from soups to self-basting poultry.” Hidden gluten: http://glutenfreecooking.about.com/od/gettingstarted/a/hiddengluten.htm
The texture is likely to be different, Feisal Kamil . I’ve not compared them..does anyone on here know?
wow- this is amazing- thanks for writing this up!!! gluten is everywhere and in foods you wouldn’t think of…
mary Zeman , I just loved the science behind this project 🙂 Also, gave me an opportunity to find out exactly what is the problem with gluten.
Rajini Rao For CNN that’s a lot of good factual information ; )
I find it really interesting and more than a little troubling how the fact that some people have a terrible problem with a particular compound causes that compound to achieve an air of being considered generally unhealthy. The lack of scientific rigor that food purists are willing to accept is something I find a bit disturbing and somewhat astounding.
Gluten is now considered evil because some people have Celiac’s disease. Though the study that Rajin links to is pretty interesting and isn’t the target of my complaint.
Though the study that Rajin links to is pretty interesting and isn’t the target of my complaint.
Anyway, I think this engineered bacteria is a really fantastic development. I’ve been waiting for biotech like this to show up for a few years now.
My sympathies, Feisal Kamil ! What’s special about Wednesday, do you know?
Do report back when you do a taste test.
Eric Hopper , most of us don’t have a problem with gluten and are only vaguely aware of it. Making bread dough involves “developing the gluten” and has a healthy, happy sound to it 🙂 I think it is a huge deal for about 1% of the population, and more a stomach irritant for another 10%. The “evilness” aura comes from the amount of hidden ingredients in processed food..that’s probably not such a good thing for all of us.
This is a friggen excellent post! Thank you SO MUCH for that. I just love love LOVE the thought through science on this.
Hehe, me too!!! John Lipscomb . I almost felt like a fond parent when I read their home page (UW site on my post) 😀
Surprised that no one here has mentioned that not all gluten is the same. Most of the problems people have are with the wheat gluten…corn and rice much less.
Can allergy-hysteria be playing a part here? Rajini Rao because Sweden is awash in self-diagnosed martyrs to gluten etc. etc. . Most are women, obese, uneducated, unemployed and unemployable looking, not for a job, but for a hand-out from the lavish welfare state. Some sit in isolated log cabins in the wilderness to shield them from their allergy to electricity, too. I have little tolerance for their allergies. Does that mean I can join the queue?
John Condliffe Actually it’s real. As a matter of fact it has gone undiagnosed for years. I believe it may be what people used to call consumption.
Gluten is a mixture of two proteins: gliadin and glutelin. It’s not found in rice, corn, quinoa, buckwheat and a bunch of other starch sources.
John Condliffe , sounds like these people have seasonal affective disorder if they are isolated in a log cabin somewhere in the winter hinterlands of Sweden, no?
Rajini Rao You’ve given an ecellent very real biochemical explanation, but do you rule out the psychosomatic component? I can vouch for the scurrilous economic one.
And yes, they are not seasonally affected but completely mentally defective.
True, there could be psychosomatic issues, misdiagnosis and inept/dishonest physicians..for just about any disorder, unfortunately. People living in Northern Europe likely see a lot more of this, whether real or imaginary, I could not say.
The biochemistry is very real, that’s all I’ll say 🙂
Rajini Rao Oh yeah, that’s right, rice doesn’t have any…I knew that, I’m having a duh day. They do call it corn gluten though or CGM and much of it is fed to livestock and used in processed foods.
Thanks, Gnotic Pasta . In the name of science, could you taste test those beers and report back? 😉
Have not heard of CGM, Jim Carver . Will look it up.
Roelf Renkema , ah yes… Ali Adelstein ‘s excellent post. I was struck by how much processed food was on the tables of most western countries shown. Personally, my diet looked a bit like the Egyptians shown.
Here’s Ali’s food post if anyone missed it: http://goo.gl/zm5AJ
Jim Carver , I found out that corn gluten (used in animal feed) is actually a misnomer. According to this link: “Unfortunately, a misuse of the term by the corn industry has become common in recent years. It has become fairly common to call corn storage proteins corn gluten. Personally, I think there is no justification for such usage. Corn may contain prolamins, as does wheat, but not gluten.” http://goo.gl/iLrO4
I suffer food allergies, including wheat so this has been so helpful
Coming back for a respectable read later Rajini Rao
Rajini Rao There is definitely a financial curve following these epidemics in Scandinavia, I hope the zombies – who “understand” everything without reading anything – won’t descend on us again, because there’s talk of corelation. Do you think they are still lurking somewhere back on Science Sunday? They’re forming long queues here is all I can say + they wear plastic bags on their heads, secure in their syndromes, hiding from the cosmic backgound radiation, with the all that dreaded mercury in their tooth-fillings generating alectromagnetic forces to stoke their well-seasoned affectations. I feel ashamed only to have suffered three brain-bangs compared to these poor souls.
Rajini Rao This is excellent, thank you. I suspect gluten may be a factor in migraine, so have eliminated it as much as possible. I am having fewer headaches, so am sticking with brown rice. I don’t really eat anything processed, so don’t think I’m getting gluten there.
Cheryl Ann MacDonald , a postdoc in my lab confessed that he would rather follow me on twitter than on G+ because my posts here are too long! 😀
I do want to write up more short posts, I used to do them more often…
Stace Irving – fyi
Thanks Rajini Rao that’s what I thought but it had been a while. I’m having a bad day with the hands and can barely type. You may remember a few years ago they had some bad “gluten” get in pet food from China and it was killing people’s pets? That’s what they were calling it and it caused a lot of confusion. Just wanted to see that point covered. Thanks again. 🙂
Haha! He is pretty smart and hard working 😉 I’ll forgive him for finding my posts boring 😀
Jim Carver , hope your hands get better…stay off the keyboard! I do recall the pet killing additives: it was melamine. The idea was that the amines released from melamine gave a high protein estimate so it was added to wheat gluten. The problem is that a product of melamine, cyanuric acid, forms toxic precipitates in the kidney. We actually did some experiments with it in yeast cells (long story).
Thanks for sharing…my mum is a celiac
Rajini Rao Thanks, it’s getting better. I still get bad days though. I need to get out my arsenal and go after it today. Epsom salts, vit. E , ultrasound etc.
Glad you found out what it was. I remember there were all these theories flying around at first. I think Mom lost her little dog because of it.
My daughters and I have Celiac disease. My youngest almost died from it, and when she was diagnosed the rest of us got tested.
Here is a link to info on gluten intolerance, celiac disease, and migraine:
http://www.celiaccentral.org/research-news/Celiac-Disease-Research/134/vobid–7959/
Congratulations on an excellent post. The hidden gluten matches the hidden lactose that is a curse on those afflicted with allergy. Thank you.
Thanks Mara Rose that’s interesting. I think I may have at least a marginal case of it myself. Wonder if that could be linked to DC? Who knows…I don’t think we understand these auto immune disorders very well.
Seems to affect the same group though…N. European descent. Same as the Viking hand.
Very interesting 🙂
Thanks for the info!
And as someone who is very sensitive to gluten I can say that the hidden gluten in foods, and contamination, are the bane of my existence and health.
I hope there will soon be a simple pill or probiotic that will break down the allergenic gluten, Julia Freewoman .
I think the yogurt is a great idea. They would probably work together and achieve a level of synergy with the probiotics.
Rajini Rao No it is not that your posts are too long….it is my time here that is becoming a bit more limited. And I want to be able to fully understand some of your posts (medical especially) for my own writings. (Some of my own posts are long :-)) BTW are we following each other on twitter and I don’t know it or what 🙂 (health psych is me)
I’m a Twitter noob, Cheryl. What’s your twitter handle,@healthpsych? I found a few Health Psychology names.
It’s usually better if you read the first 1000 or so comments in order to find out if your topic has already been covered. 😉
Send me your link https://twitter.com/HPofSD I am more acquainted with twitter for my own personal use (it is wonderful for news….the best yet IMO) Google is much more relationship oriented and is the best IMO 🙂
Here’s a good article written in layman’s terms on the difference between food intolerance and food allergy and talks about celiac disease. This condition is not really an allergy.
http://www.food.gov.uk/multimedia/pdfs/allergyfactsheettwo.pdf
I can get to the homepage http://www.food.gov.uk/ but not to the pdf link (I get a 404 page not found error).
Hm, Not sure why…it loads for me. I tested it on the link I posted.
The pdf loads fine for me, too.
OK, may be it was a time out issue. It loaded on the third attempt. Thanks!
Seems the criteria here is the time frame. If you are allergic to a substance, you will get a reaction right away. In the case of intolerance, depending on severity you may not be aware of it. It may seem like slitting hairs since an intolerance likely means you are allergic to it to some degree. But it’s still an important distinction I think.
Rajini Rao Thanks for the post.
Just curious: does this project impact your perception on de novo protein structure determination and (re)design? Are the tools/methodology looking more useful to your eye (i.e. less gimmicky)?
Vladimir Vigdorovich , you know, I really was thinking of you when I wrote this post? Mea culpa, yes..I think this definitely vindicates and validates the protein design concept for me. BTW, David Baker is co-author on the paper 🙂
It is definitely fun to see an obscure and weird-sounding methodology break through with such a potentially far-reaching impact for so many people. Gotta love science!
As for DB, I wouldn’t at all be surprised to learn that he invested some time to mentor this group. I remember being inspired by his talks and personality more than a decade ago.
That’s why I put in a plug for basic research. People may wonder about the usefulness of cloning, purifying and solving the crystal structure of a protease from a thermoacidophile. But it was that obscure structure that was the basis of a design that could have such high medical impact.
Here is the thread from a couple weeks back …
https://plus.google.com/u/0/113677817925719341659/posts/aggR98QvWgE
The source article makes a bold statement that seems worth exploration: “today’s wheat is a far cry from what it was 50 years ago.Back in the 1950s, scientists began cross-breeding wheat to make it hardier, shorter, and better-growing. This work, which was the basis for the Green Revolution — and one that won U.S. plant scientist Norman Borlaug the Nobel Prize — introduced some compounds to wheat that aren’t entirely human friendly.”
Good
Another possible factor in the increase in celiac disease and other autoimmune disorders is the “hygiene hypothesis” that our immune systems develop differently now that we live in environments that exclude many kinds of germs and parasites. http://www.celiac.nih.gov/prevalence.aspx
Since rolling back to an environment with more bacteria and parasites is not exactly a good idea (putting it lightly), and modern wheat has a lot of advantages, approaches such as this enzyme and the vaccine currently in trial in Australia are going to be key.
Gretchen S. the Hygiene hypothesis is the reasoning behind introducing relatively benign parasites back into our bodies in a controlled way..I had a post on this some time back.
joe breskin , thanks for the link. From what I’ve read so far it appears that (1) we are now exposed to more wheat products and gluten than in historic times when we also consumed other grains like millet, quinoa, etc. There’s also the issue of gluten used as an additive in processed foods. (2) the gluten content in wheat is increasing according to this article in the link below. Basically, this is due to the increased ploidy (chromosomal copies) of newer varieties of wheat which would increase the number of gene copies of gliadin.
“Different types of wheat have different numbers of chromosomes, and some studies show that the older wheats, with fewer chromosomes, tend to have lower levels of gliadins, the type of gluten proteins that seem to cause most sensitivities.
Einkorn, the oldest known type of wheat in our current food supply, has just 14 chromosomes, and is called a diploid wheat. Durum wheat (the kind most often used for pasta) and emmer are tetraploid wheats, with 28 chromosomes. Common wheat (used for most everything) and spelt have 42 chromosomes and are known as hexaploid wheats. Research shows that different tetraploid and hexaploid wheat varieties differ widely in gliadin levels, and it’s possible to select “individual genotypes with less Celiac Disease-immunogenic potential.”
http://wholegrainscouncil.org/newsroom/blog/2012/01/research-sheds-light-on-gluten-issues
Are the numbers really going up or is it better screening and population growth? Guess that’s hard to say.
Gretchen S. I think you can go too far with the hygiene hypothesis. It has some merit in certain cases but I don’t think this is one of them. I think the other possibilities in that article are likely if you have a genetic predisposition.
Genes linked to gluten sensitivity are variants of antigen presenting HLA-DQ genes. I had no idea the whole spectrum of gluten sensitivity was so complicated http://en.wikipedia.org/wiki/Gluten_sensitivity
hallo how r u
Rajini Rao yes, that ploidy issue was part the discussion I have been having with my recently-deceased baker-friend Frank D’Amore over the past couple of decades.
And Jim Carver yes, I really think it (whatever the problem turns out to be) is increasing, beyond what I can easily attribute to fad or awareness. In my post of this story on FB (a petri-dish community of people that I actually know in meatspace) I got nailed repeatedly in comments for opening with the observation that this was something that was affecting a growing number of my female friends. Which is fact. But my population is pre-screened by the simple fact that – aside from myself – I almost never cook for males
And a substantial number of my male friends noted that they had been shifting away from wheat. The pool of women for whom I tend to cook meals ranges from around 27 to around 67 and most of them are now avoiding wheat … Which led me to ponder what ELSE could be going on. What combination of factors might be involved.
joe breskin Fair enough, I just wanted to bring that up to make sure. I knew it certainly appears that it has. Looks like the rate of gluten intolerance has been increasing also, not just celiac sprue. (I’m even learning new words:)
Looks like people’s reactions are quite different also. I was aware that there is a large variability in reactions but it’s really complicated. One comment on a forum by a lady said she could eat spelt but nothing else in the group. Interesting, well I have to go check my rice and beans. 😉 Really, it’s true.
Rajini Rao I’ll have to look for that post — I’ve seen some studies on using worms to control other autoimmune disorders, because they secrete immune suppressants. In general it sounds to me like a drastic approach.
Another very drastic approach is bone marrow transplant; there have been cases of transplant both curing and transmitting some autoimmune disorders such as celiac disease. (Relatively benign autoimmune disorders such as celiac disease do not preclude becoming a donor, because they are far less harmful than the diseases the transplant is being used to treat.) http://pediatrics.aappublications.org/content/120/4/e1120.full (for a case of remission) http://www.ncbi.nlm.nih.gov/pubmed/9337064?dopt=Abstract (for a case of transmission)
Jim Carver I don’t think it’s out of line to look into the changes in bacterial environment as a factor for the rise of incidence, given that it is theorized that there was positive selection pressure for the gene in question as protection against bacterial infection. http://www.ncbi.nlm.nih.gov/pubmed/20560212 That line of investigation may lead to an approach that could prevent onset in genetically susceptible people. I would like to see a study following up the one that screened for antibodies in the blood samples from the 50s, that screens for incidence of the genes, to see if there is a different percentage in onset. But yes, better diagnostic techniques and understanding are definitely another major factor, as could be the changes in modern wheat and the way we use it.
One note about screening for those who suspect that they might have celiac disease or gluten sensitivity: the presence of the genes does not indicate presence of the disease. Celiac disease can activate at any time in life in a genetically susceptible person. The antibodies that indicate presence of the disease can reduce to inconclusive levels in as little as two weeks, so while it can be useful to go on an elimination diet to see if symptoms are relieved, it is very important to be actively eating gluten prior to the blood test or the biopsy. Otherwise those tests can come up negative. Since the gluten-free diet required to keep full blown celiac disease at bay must be very rigorous, having a clear diagnosis can be key for keeping people with active celiac disease strictly gluten-free. Self-diagnosis is just not a good idea, and a gluten-free diet is not a weight loss diet. (For people with celiac with malabsorption issues, as is common, it can actually be a weight gain diet.)
This may be all redundant and stuff, but I wonder if enteric coating of the Sphingomonas capsulata would protect its transit through the stomach. Bet they tried it…
Hopefully the proven treatment won’t involve a fecal transplant.
Jim Carver I do not think anyone I know actually has anything clinically diagnoseable as Sprue, but I know at least a dozen people from that pretty small pool who I actually eat with fairly regularly, who have “discovered” over the past year or two that they do a lot better w/o wheat. To the point that I currently have a substantial stock of rice-based pasta in the kitchen.
FWIW, I have run my own body as an experimental station for the past 45 years, and actually had a food-based cult back in the early 70’s based on dogma amalgamated from other dogmas that I made up to give people something “safe” to talk about.
But I do not believe people are very good at figuring out what is important in their diets My mom did clinical nutrition focused in particular on Zinc deficiency and fetal alcohol syndrome. And one winter in the early 80’s I got to help her design a large scale experiment that traded chronic alcoholic ‘street people’ a warm bed and free food for participation in a drying out program, and the experiment involved a menu with a spectacular range of food choices, with the unstated goal of comparing their actual blood values with what they wanted to eat or what their bodies felt like they needed to eat …
For a few years in the mid-90’s I ran my own body GF just to see what would happen, but found the resultws inconclusive and the inconvenience unjustifiable. At the moment, at age 65, I am beginning to believe that the unfair head start of privilege (good genetics and good health) that I had on board when I left the launch pad probably carried me this far effectively regardless of the experimental diets (macrobiotic, paleo, hunter gatherer, etc.) that I’ve explored.
David Archer , the bacteria might survive but the enzyme would not. The problem is that the Sphingomonas enzyme works optimally at pH 7 and it is not that easy to engineer an acidic optimum in a protein.
Rajini Rao Hence why they started with an acid-tolerant enzyme in this case?
joe breskin Wow that’s pretty awesome. I’ve often thought I got such a good head start also that not much would hurt me too badly. We always had good food and a few supplemental vitamins. I knew we were different in our family because I couldn’t eat the food in the school cafeteria and those other kids would just eat it up. Boy, you talk about some glutinous gravy on that fried steak.
Hmm. I see, Rajini Rao. Not a lot of reliable pH 7.0 in the human digestive tract, I’d expect (and hope!).
Gretchen S. , exactly! The enzymes from bacteria that live in hot/acid springs are already optimized to work in acid conditions. Starting there, they were able to tinker with the substrate binding pocket..they made just seven changes in all but had to check them in various combinations.
David Archer , the stomach is highly acidic..close to pH 2 actually. That’s where the initial protein digestion begins. The intestine is neutral to alkaline. The digestion and absorption happen there.
Rajini Rao I find it very exciting that they were able to make use of the protein folding simulation in this research. It seems like we can cut out a lot of dead ends by using simulations.
I found your post on whipworms and it’s very interesting, thanks! I’m sorry I missed that earlier. I had heard about the work with hookworms but they have the issue that they are not exactly benign.
Here’s a link to the worm post for anyone who wants to refer to it: https://plus.google.com/114601143134471609087/posts/epN3K2VZzQ5
Ok I am home, nice and cozy with Tessie the Jack Russell plastered to my body like a band-aide. 🙂
I really like the so much win section and using video games to do the research. Video games IMO have a bad reputation and tend to get blamed for many problems not yet proven to be true.
I was not aware of the link to cancer ….but it makes sense http://www.celiaccentral.org/Celiac-Disease/Related-Diseases/Intestinal-Cancer/46/
Aside from using diet as a prevention for cancer with those diagnosed celiac disease …. does science have any other treatments?
Cheryl Ann MacDonald , have you ever posted a pix of Tessie? She sounds cute 🙂
Yes, the Fold-It part was great, expecially as these were all young undergrads. They must have had fun with it.
The cancer connection comes from inflammation. I believe the same is true for gastric ulcers and gastric cancers. Unfortunately, the only current treatment for celiac is to keep away from gluten.
Gretchen S. , I was dubious about Fold-It earlier but this paper really convinced me that it is great for virtual design of new functions. Glad you liked the worm post..fascinating isn’t it? Now they have fecal transplants too, replacing bacteria by transferring stool from a healthy person :O
I did find this:
“Very high concentrations of prolyl-endopeptidase have been shown in biopsy studies to reduce the amount of immunostimulatory gliadin peptides (both innate immune and T cell activating) reaching the mucosa.116 Whether these therapies will work as dietary supplements awaits human in vivo trials. Such supplements may be able to prevent inadvertent low level gluten exposure even if a full gluten containing diet cannot be achieved.”
http://gut.bmj.com/content/55/7/1037.full
Cheryl Ann MacDonald The diet reduces risk of colon cancer for those with celiac disease, and can prevent anemia and osteoporosis in those who are experiencing malabsorption from the disease. Symptomatic celiac disease is also linked to infertility in some women. So is some lactose intolerance (because the disease attacks the part of the small intestine which digests lactose in normally lactose tolerant individuals), GERD, and IBS. The symptoms go completely into remission on a fully gluten-free diet, usually taking six months for the symptoms to clear, though it can take longer.
Right now there is no known cure for celiac disease other than a possibility of bone marrow transplant, but only people with far more serious illnesses such as leukemia receive those transplants; it is not done simply for something like celiac disease for reasons that are probably obvious. This makes me think that stem cell therapy would be another possible approach, if stem cells become available enough to treat less serious diseases.
There is a vaccine based on immunotherapy currently undergoing trials in Australia. Last I read it will be out of human trials in 2017. I don’t know what that means for people in the US or if the FDA is looking at it yet.
Jim Carver An effective enzyme would be really useful for celiacs to be able to dine out, even if it isn’t able to handle large quantities. Even if it doesn’t protect against eating, say, a wheat bread sandwich, it could protect against the low levels of gluten that can cause a reaction if, say, the person who made your gluten-free sandwich used the same knife on wheat bread first. Cross-contamination is a big issue because the concentrations that can cause a reaction are so low.
Wonderful information, thanks Gretchen S. .
The review you linked to, Jim Carver , looks like a great read! They bring up the health impact on osteoporosis, cancer and infertility that were mentioned above and much more.
Rajini Rao That’s a good one. I haven’t read it all yet and some of it just over my head. 😉
Excelente información, gracias. Yo tengo perfil celiaco
Excellent information, thanks. I have a profile celiac
Thank you Gretchen S. this was very important information to share…ahh a difficult one to apply being that it is all diet/food related.
Tony Kocurko It was covered but you probably missed it. It was linked in joe breskin ‘s post…sort of.
Okay I found the research article for the discovered form of prolyl endoprotease and it’s made from Aspergillus niger being optimal at 4-5 pH and stable down to 2.
http://www.ncbi.nlm.nih.gov/pubmed/16690904
Ooh, I wish my wheat allergy caused weight loss! [/inappropriate comment]
Interesting post!! This is something we are seeing in dogs too.. Wheat, barley and gliadin contaminated oats are common ingredients in dog foods..
The gliadin protein (and similar lectin proteins in other foods like corn, soy and dairy) are known to cause autoimmune diseases as well — like MS, Lupus, rheumatoid arthritis, diabetes etc. In fact, lectins (and lectin like proteins) can bind with insulin receptors resulting in insulin resistance, weight gain etc.
The body protects us from lectins by making antibiody IgA and mucin which bind with the lectin and allow it to pass harmlessly. Probiotics help stimulate IgA. However with repetitive consumption of the lectin foods that we are intolerant of our protective devices wear down and symptoms appear (sometimes months and years later).
Lectins from certain foods can also cause the little hairs in the intestines (the villi) to die back — referred to as villous atrophy. Malnutrition (despite the quality of the diet eaten) is the end result.
The lectin in the casein protein in dairy have caused this same damage to me — malnutrition (along with all the symptoms of that — iodine deficency hypothyroid as an example), insulin resistance etc..
In dogs it can manifest as IBD, IBS, colitis, diarrhea/vomitting, and autoimmune diseases..
Sorry for the ultra long post…
Great information, thank you Shawna W ! I’d not considered the effect of glutens on pets. Perhaps the best bet (for those of us without gluten allergy) is to diversify our food sources, eat a lot of different grains and legumes so that no one food dominates and overwhelms our ability to handle it.
Absolutely Rajini Rao!! It won’t prevent the intolerance but it will allow the body recovery time between exposures and therefore fewer to no symptoms..
It is best not to over consume any one food as many different foods have lectins that can be damaging to different people — like cucumbers, strawberries, kidney beans (all legumes including green beans and peas), dairy etc. The foods most likely to cause issues for the most people (but not in every person) and/or being the most damaging are grains (with rice seeming to be more tolerable), legumes, dairy, corn and white potato (actually any of the nightshade plants).
I was making ham and bean soup and forgot to poor off the soaking water from the pinto beans. I was fine but we learned that evening from the VIOLENT symtoms my hubby experienced that he is VERY reactive to the lectins in pinto beans :(… He has refused to eat them since — that was about 10 years ago… Soaking legumes (AND pooring off the soaking water) helps remove many (but not all) of the lectins in legumes.
Sprouting grains is a good way to deactivate the lectins in grains.. However there are some still remaining so those with celiac will likely still have issue. Those with gluten intolerance may get by..
Great post!! This is a topic that definitely needs more awareness… 🙂
A good reminder to toss out the soaking water from beans and legumes, Shawna W ! I didn’t know about sprouting grains reducing some of the allergens..they are supposed to be more nutritious after sprouting, although I’ve not looked into the science behind it. Certainly, they cook up faster, so that’s a plus.
I had an old post on toxicity from common foods, nuts and veggies: https://plus.google.com/u/0/114601143134471609087/posts/59M7JDwUymF
There’s some good data on the increased nutrient profile of sprouted grains. Some of the best reasons for sprouting — 1. sprouting deactivates the antinutrients in grains (like phytic acid). Phytic acid binds with minerals and prevents their absorption. Diets high in rice, as an example, can cause a zinc deficiency (there’s science papers on google scholar regarding this) 2. sprouting deactives the enzyme inhibitors (appropriate cooking will do this too however) 3. the grain becomes a “living” food with active enzymes 4. Sprouting also increases the protein and vitamins. The data can be found on http://www.nutritiondata.com (compare barley to sprouted barley).
Off to read your post on toxins in foods..
Some people have become fearful of eating sprouts because of outbreaks of pathogens. It’s really good food though. I think I know how to do it using O3 and UV if someone was worried.
I think if you look at the foods most people like, they are almost all acid forming. All meats, most grains and pasta….everything processed with flour. It’s easier to list the alkaline forming ones: like raw vegetables, fruit w/o added sugar, honey and lemon.
So anyway, I think a big part of this is acidosis and when you stop eating these things like bread and meat, your pH goes up where it should be. I read some reports of stomach pH in some people was ,< 1.0. So I think it's likely the whole system is too low to operate the way is should. Btw, coconuts and soured milk products are low-level alkaline forming…but milk isn't, it's high- level acid forming. Milk, meat and bread, sounds like the American McD lunch.
True about contamination. Those concerned can actually purchase sprouting kits and do their own. Another fabulous way to increase the nutrient profile of foods is to ferment (or culture) them.. Fermenting (home fermenting) increases good bacteria/yeast, vitamins like B and K, enzymes and preserves the food (if stored in the fridge after fermented) for up to 3 or 4 months.
I attended a very interesting 2 day seminar in 2009 titled “Back to School for Doctors” (my father is a Naturopath and I attended with him). The presenter had a very interesting view on acid/alkaline. He felt that it was more important to eat healthy food (organically raised and fed as one example) over foods that are alkalanizing or acidifying. Made sense — example, the stomach makes hydrochloric acid to digest protein (be it animal or plant based protein). When the pH gets acidic enough it activates the enzyme pepsin (also released in the stomach) which digests the protein. If the stomach doesn’t reach proper pH then pepsin isn’t activated and the proteins may not be thoroughly digested. Additionally, acids in the colon (such as butyric and lactic acids) help prevent cancer and keep the bad bacteria that get in (possibly through contaminated foods) in check. It is the good gut bacteria that create the two acids by consuming the prebiotic foods they like (starches and fibers).
I think the quality of the meat has more relevance to our health than being meat free.. It is well studied that cows eating grains, soy and corn produce meats with more saturated fat and less omega 3 fats than grass finished cows. When you factor in that some, if not most, of the grains etc fed to the cows are genetically modified it introduces a whole new dynamic..
Great and stimulating converstations!!!!
Shawna W Certainly no strangers to fermentations around here 🙂 I’ve always got a pot of yogurt or cheese or even some peppers going. I like to cook a lot in the winter too so it’s heats the kitchen. Can’t stand cold kitchens…it’s a terrible waste of space…
Have you seen Rajini Rao’s blog? It’s really good:
https://madamescientist.wordpress.com/
Homemade yogurt is wonderful! As are fermented batters and doughs 🙂
They are…I’m going to make miso and maybe natto pretty soon if I ever get off my can and order the culture. I always liked doing these things in winter and controlling the heat. It’s a lot easier than trying to cool it if things get too warm. That’s really true with alcohol ferments. A large batch generates a fair amount of heat at the start.
I’ve talked about this before but I use the food dehydrator for the yogurt. It holds a nice 110F and you could fit about three gal. in there. I don’t make that much at a time though. Usually 3 liters is good.
Rajini Rao Do not worry about the length of the post. We like your posts. We can read as much as you can write…
Whoever said they are long can be ignored 🙂
mandar khadilkar I had forgotten about that, it’s been so long ago…:)…kidding. This thread has a lot of good stuff in it.
How about short posts for weekdays and longer ones for the weekend? That would make it easy on both you and me 😉
Haha…well I haven’t given up yet:
“Oral supplementation with enzymes that can cut gluten has been suggested as a potential treatment modality for coeliac disease. In the present study the investigators wish to determine if co-administration of such an enzyme, a prolyl endoprotease derived from the food grade organism Aspergillis niger (AN-PEP), is capable of detoxifying 8 grams of gluten in a commercial food product.”
http://clinicaltrials.gov/ct2/show/NCT00810654
Any idea on how to read it, or do I have to search for it?
Jim Carver , that was an interesting find! It looks like the clinical trial was done in the Netherlands and was concluded in 2009. However, I’m not finding any publications on it. The study lead, CJ Mulder, has several recent publications on gluten intolerance, but nothing on the outcome of a clinical trial. I guess it was not successful.
The A. niger enzyme is not as specific as the one these kids have designed. Because of their selection process, they came up with a highly active enzyme. Also, proteins from archaebacteria, such as thermoacidophiles, are highly stable so I think they picked a winner. It looks like the senior authors of the study have a start-up company and perhaps someone will license the enzyme from them and run a clinical trial in the future.
Rajini Rao Thanks, I couldn’t find anything either. It’s like it went into a black hole..I’d still like to see the numbers. Maybe it did totally fail because it wasn’t specific enough and got swamped. Or maybe there was a problem with delivery…idk, I’m just guessing.
Well anyway, we got this topic pretty much wupped! (for now) I learned and re-learned a lot. I added an excerpt from the article into the wiki on celiac. So there’s a lot for people to access these days. When I first started talking about it years ago, hardly anyone had ever heard of it.
Shawna W Another issue with dog food is that it is very hard to find dog food that is made to the same standards as people food. I don’t like to handle food containing gluten, because of the possibility that I’ll then cross-contaminate my own food (especially as we feed them in the kitchen) and on making inquiries have found that most brands are on shared lines and they will not guarantee against cross-contamination. Plus makers of so-called “grain free” brands who think that barley doesn’t contain gluten and that barley isn’t a grain. (It is very much a grain, and while it contains much less gluten than wheat, it very much contains gluten. And oats are almost always contaminated with gluten unless they are grown and processed in dedicated gluten free fields/facilities.)
We have had three dogs that had various kinds of grain intolerances, one to corn and two to wheat, quite often to the point of constant vomiting if someone slipped them a milk bone, poor critters; the latest has stopped having bright pink eyes that wept crud and horrendous dandruff after we put him onto grain-free food. We normally slowly taper a dog off the old diet upon adoption but he was so clearly having allergic reactions when we adopted him that we not only switched his food immediately, we put him on benadryl (on our vet’s suggestion) during the transition. We do treat the food as potentially cross-contaminated, so I scrub my hands after feeding him or if he licks me.
Someone else got some of those home ELISA kits for testing for gluten and ran it across 10+ brands of supposedly gluten-free dog food and found that only one brand met human standards of gluten-free; most were more than 100ppm which is more than enough to cause a serious reaction in a human. I guess the moral of the story is that if you’re celiac, treat even “gluten-free” dog food as if you are handling foods containing gluten, and wash your hands thoroughly. For celiacs with DH (which can trigger on skin contact) I suspect they’d want to use gloves or formulate their own dog food.
Hi Gretchen S. , I wanted to tag you on this other post on parasites, inflammation and obesity but couldn’t get your name to come up. But here you are, so check out: http://goo.gl/Kyl3X
Rajini Rao Oh, that’s very interesting! I wonder if, in a generation or so, it will be matter-of-course for people to be deliberately infected with strains of parasites that have been toned down so they are less of a danger/burden but still release all of the compounds that we find helpful. In much the same way that it is a good idea now to re-seed with probiotic cultures after a course of antibiotics.
That’s an interesting comparison with probiotics! I’m not a parasite expert, but it sounds hopeful to me.
Sounds pretty weird ;~)
“Chronic inflammatory responses found in most autoimmune diseases and metabolic diseases exhibit common characteristic processes where macrophages are initially activated and interferon (IFN)-γ-producing type-1 T helper cells subsequently stimulate macrophages to release more inflammatory cytokines. Together with our previous findings that CWE prevented anti-CD3-stimulated T cells from secreting IFN-γ, our current study clearly shows that CWE is able to interfere with the chronic activation of macrophages [16].”
CWE= Cinnamon bark Water Extract
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3533872/
I did see Suzanne Catty ‘s post on this and shared it via ScienceSunday yesterday 🙂
Chad Morey
FYI to anyone interested in this topic. Jim Carver was kind enough to compile all the links relating to gluten intolerance that were discussed in this thread here: http://goo.gl/Jkmh8
I’m thinking the wheat hasn’t changed as much as most of us have. Was there any less gluten in the wheat grown 1000 years ago? I see nothing to indicate that. I do see a massive shift in the way and kinds of foods people eat in the past 100 and especially 50 years.
Very kind of you to mention that Rajini Rao 🙂
Gretchen S. yes, I agree completely.. One of my dogs has an issue with gluten. I have been feeding my dogs raw for about 7 years. It’s a bit more time consuming and can be a bit pricier but its nice to have complete control over what they are eating..
Looks like this thread has gone to the dogs! ;D
Haha! 🙂
I had one more thing to mention about yogurt, since we were talking about probiotics and yogurt: I have successfully cultured two of the ‘proprietary’ formulas. The one I remember was Activia. It came out the same consistency and tasted the same. So if it looks like a duck…
For the price of one of those small tins (how archaic), you can make a whole gallon of the stuff.
The other one I can’t think of the name, but it worked also. Use the same technique as regular yogurt and I wouldn’t re-culture. Maybe once is okay. I usually don’t re-culture anyway; of course if you use dry culture that’s a different story. I doubt you could get these that way, I think they’re patented.
Excelent information !
very useful info…
Thanks!
Still hope for my wife then.
I hope so, Robert Ferentz . If lactose intolerance can be so easily handled by pretreating milk, we should be able to do the same with gluten.
Update: I just came across a news story of early success on clinical trials using hookworm therapy to treat celiac disease/gluten allergy: http://www.abc.net.au/news/2014-09-25/hookworms-coeliac-disease-research/5768732
Here’s an old post that explains how hookworm infection helps our immune system: As the Worm Turns https://plus.google.com/u/0/+RajiniRao/posts/epN3K2VZzQ5
The idea is that our immune response is tamped down and this helps relieve the reaction to gluten. Here is an open access publication from the authors of the clinical trial: http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0024092
Hope this helps!
Haha, apparently they did find volunteers so I’m guessing that there are some celiac sufferers out there willing to deal with the grossness factor.
Gnotic Pasta I didn’t pick up on that! I assumed the patients swallowed a drink laced with eggs. I’m curious if people with celiac would seriously consider this approach.
Especially if it’s hard to control with diet. It’s more than an inconvenience, since prolonged inflammation increases the risk of cancer.
The way to cajole people into this would be to give them a nice, comprehensive overview of the average human microbiome. With numbers of species, population sizes, comparative mass, and best of all, pictures (e.g. Demodex). Then once you’ve reduced the squeamish to trembling tears of revulsion, “what’s one more?” would be pretty anticlimactic. No?
There’s research in progress towards modifying the wheat genome to not produce gliadin – that’s great, but the logistics, cost and availability factor are a much more complicated proposition for the average sufferer, certainly for the near term (and that’s assuming the project bears … fruit, so to speak).
David Archer there’s evidence that gliadin content has been enriched over the past decades of selection and cultivating wheat varietals. It makes sense to reverse engineer this. Good to hear of this research.
Rajini Rao I’ve looked into hookworms but it seems to be a tricky balance…. It’s a short distance from a thereputic population to anemia (which people with celiac disease are already at risk for.) Some people are using them very off label to control severe allergies or really hard to live with autoimmunes like Crohn’s. There has been some research in using variants from pigs that don’t multiply as well in humans…. They still secrete immunosuppressants but don’t make themselves quite as deleteriously at home.
Personally I think the vaccine approach going through approval in Australia has a better chance of being a therapy I’d actually try, and I’m pretty impaired by autoimmunes. But if my allergies were severe to the point of regular ER visits, hookworms would start to sound interesting.
Thanks for looking into this for us, Gretchen S. . Glad that you mentioned potential anemia as a complication of hookworm therapy. The use of pig-infecting variants sounds like a clever compromise. Let’s hope that more breakthroughs are forthcoming..we should be able to lick this thing!
Rajini Rao I actually was solicited for a study for the deactivating enzyme pills, which are a good idea (they don’t make it safe to eat a pasta dinner, but they make it much safer to eat out since they can handle a gram or so) but I had to regretfully pass…. I get really wretchedly sick for months and being in the control group would suck too hard.
Rajini Rao I think the parasite research is really interesting because they are a dynamic system – immunosuppressants in general are very dangerous for all of the obvious reasons, but something like hookworms that has evolved to not kill the host could become effectively a trickle-dose delivery system that acts in concert with the host…. It’s a more delicate touch. But we need to keep them from multiplying unchecked and from spreading to humans that don’t need them! While modern bathrooms really help against worms, my partner was justifiably concerned by my research.
Another concern, cross infection! Thanks for the perspectives, Gretchen S. .
http://www.jacionline.org/article/S0091-6749(14)01010-0/abstract — I tracked down the abstract of the pilot study; it’s very interesting! The reporting on this is very enthusiastic yet manages to imply that only symptoms are used for tracking, which made me concerned (while they likely selected patients who tend to present obvious symptoms, celiac disease can be relatively asymptomatic while still doing a lot of damage to the small intestine), but when I looked at the abstract it looks like they also do serological and other tests to measure tolerance. Better yet, they are trying to isolate the secreted protein, because ground-up hookworm powdered secretions are (amazingly) a lot more appealing than having a live colony of hookworms. The 8 remaining participants of the pilot study all opted to keep their colonies, though! And none of the dropouts were related to either the worms or celiac symptoms, according to the reporting. I believe this is a better dropout rate than the study that measured reactions to gluten-free oats.
Between this, the vaccine in the works in Australia, and the enzymatic breakdown pill currently in testing, it seems very likely there’s going to be some reasonable control measures or even a cure available for celiac disease in my lifetime. Science is wonderful!
Thanks for tracking down the study, Gretchen S. ! Hookworm powder is so much more appealing, I agree! Here’s hoping that one or more of these promising leads becomes a routine adaptation soon.