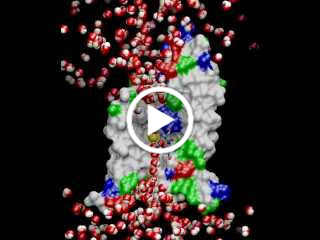
Occupy The Cell! 99% of the molecules in our cells are water, about 70% by weight .
“Water is life’s mater and matrix, mother and medium. There is no life without water”, so said Hungarian biochemist and Nobel laureate Albert Szent-Gyorgyi (1893-1986)
Water enters and exits the cell through water channels known as Aquaporins, embedded in the hydrophobic lipid membrane.
Aquaporins move water at the rate of 1/ns or 1 billion water molecules per second!
The animation shows water molecules squeezing through a narrow pore in a single file. One of them is labeled in yellow so you can watch its progress through the 10 ns simulation. Notice that the oxygen atoms (red) face the channel center, flipping in orientation in the middle of the pore. This is thought to break the chain of hydrogen bonds and prevent protons from tunneling through, which would destroy proton gradients and mess up cellular pH. (See Grotthuss mechanism: http://en.wikipedia.org/wiki/Grotthuss_mechanism)
Discovered by Hopkins scientist Peter Agre by accident (he was searching for the Rh antigen on blood cells); my colleague Bill Guggino tells the story of putting the unknown protein into frog eggs ( Xenopus oocytes, used because of their large size) to measure their function..but water rushing into the eggs made them explode, spattering yolk all over the microscope lens! Agre was awarded the Chemistry Nobel in 2003.
This molecular dynamics simulation is by Emad Tajkhorshid (http://csbmb.beckman.illinois.edu/) and is featured on the Nobel web site.

Water – let’s keep it real!
Fascinating!!
Exploding frog eggs?… I bet the mythbusters would be interested. :p
Very interesting indeed, amazing dynamics to block and facilitate tunneling.
Koen De Paus , with reference to my earlier comment on valinomycin, once again we see that a simple bacterial antibiotic, gramicidin (http://en.wikipedia.org/wiki/Gramicidin), can efficiently move water through its pore, but cannot keep out the ions. As a result, gramicidin kills cells. Aquaporin is much more complicated because it is selective to water, and not much else. 🙂
If only we were that simple! We would have us all figured out in no time… We would also be bacteria but… hmz, nevermind. 😉
Haha, yes, I was trying to make the point over a number of posts that there is a price to be paid in complexity of design (also in cellular cost of making a big protein) for increased functional sophistication! Bare bones function is really easy to code but not good enough for us.
Amazing … can it selectively transport water in one direction or does the water move in response to whatever the concentration gradient across the membrane happens to be?
Matt Kuenzel , water can only go down its gradient, whichever direction that may be (another way to think about it..water goes from low salt to high salt concentrations, since the concentration of water is inversely related to solute). Aquaporins are water channels. If they moved things uphill, they would be called pumps and would need a source of energy (ATP, light, etc.). For a long time, scientists did not believe that water needed channels to cross membranes..they thought that the molecules just got through somehow. But it turns out that nature was too impatient to wait for some low rate of permeability and has evolved a specialized, super fast channel 🙂
Ok I see the difference. Interesting. When I studied biology I recall reading that the cell membrane was permeable to small molecules like water.
Matt Kuenzel , that’s what we were all taught. Perhaps, permeable at some low level of diffusion, but not at the rate of a billion waters per second. Some tissues depend on rapid water movement.
Another remarkable feature is the fact that the channel is the proper size to admit only single water molecules: large enough to admit water while too small to allow more than one at a time. And the mechanism in the center that must be involved in the orientation flip.
Ah, but Matt Kuenzel , have you wondered how a channel can be selective for something like say a K+ ion, but not allow a smaller ion of the same charge, like Na+ or Li+ to go through? Surely, the smaller ion can slip through..but it does not. Any ideas?? 🙂
+Мултановский Вадим , van der Waal interactions are important..true. I was hinting at the hydration shell of an ion in water. There is an energy cost to removing it that has to be replaced by a favorable interaction with the protein channel (van der Waals or ionic interactions).
Yes, that’s right, Vadim (am I right with your name?). Because of the poor fit inside the K channel, the energy cost of dehydration is too great for a smaller ion like Li or Na and so it remains hydrated, in water. 🙂
Yes 🙂 Pronounced rudge-nee (no need to say all those i’s). Thanks 🙂
Rajini Rao I’ve wondered about the selectivity question … so it’s related to the hydration shell … And think how crucial it is … the Na and K gradients are crucial for lots of stuff like neuron spiking, isn’t that true?
Matt Kuenzel , the selectivity for Na and K is very important. Their gradients (about 10 fold each) are opposite: inside the cell, more K, whereas outside the cell more Na. Action potentials are caused first by Na channels opening (Na comes into the cell) followed by K channels opening (K goes out of the cell). As a result, there is a spike in membrane potential (in mV) going from negative to positive and back to negative. This wave form can travel down the nerve as an electrical signal. The gradients also do other kinds of “work” like moving sugars and other metabolites or for water movement (the water follows the ions, being osmotically obliged).
Rajini ji r u a chemistry teacher i hate chemistry
thank you you r great on giving information
as i h,ve studied medical jurisprudence so this type of informations r very impressing for me
Alexander Macias , I’m a curious creature and science nerd at heart, just like you. Cheers 🙂
could u plz tell me about the types of cell cultures used in virology?
Hey, Erum Atif , I don’t know. You should be able to look it up, good luck.
thanks
nice good luck
I have a question for you: you use the NAMD to improve your simulation? if yes how many time step you have set to produce that film? for my little experience I think that are about one million ..
the protein and the membrane are fixed during your simulation? I trying to reproduced, in a very very little scale, your work for an exam at University .. thank’s for the answear .. Sergio
P.S. is a very nice work 🙂
Sergio Tarantino , the simulation was done by Dr. Emad Tajkhorshid and his lab web page is: http://csbmb.beckman.illinois.edu/. You can also check out his publications on PubMed. He uses a method called steered molecular dynamics. I believe this is a 10 nanosecond simulation (don’t know the time step?) and it does seem that the protein and membrane were fixed for this..they also do all atom simulations. Hope this helps, good luck with your exam, Sergio!